Research

Although the majority of the human genome is actively transcribed, only 2% of it codes for proteins. This indicates that a large number of transcripts are non-coding RNAs (ncRNAs). While the biology of a few ncRNAs is already fairly well established, the mechanistic role of the majority of ncRNAs remains elusive. To address this lack of knowledge, my lab combines biochemistry, functional genomics and structural biology to study the molecular mode of action of non-coding RNAs that are biomedically highly relevant and whose mechanism of action has, so far, not been elucidated. Understanding the role of non-coding RNAs has profound implications for the treatment of human disease. This can be seen in the first ever RNA interference-based drug approved in 2018 and in the growing success of treating serious human diseases with antisense oligonucleotides. As a messenger RNA (mRNA)-based vaccine enabled the world to curb the thread posed by SARS-CoV-2, the COVID-19 crisis marks yet another instance, where a detailed understanding of the life cycle of mRNAs - including their regulation by ncRNAs - is of great importance. These examples demonstrate that tackling the function of non-coding RNAs using basic biomedical science benefits human health and aids in designing and improving nucleic acid-based drugs.
Specifically, our work addresses the following questions:
- We investigate how piRNAs (Piwi-interacting RNAs) influence the regeneration of planarian flatworms by regulating mRNA turnover and by directing epigenetic changes in their stem cells.
We study how eRNAs (enhancer RNAs) direct promoter-proximal pausing of RNA polymerase II (Pol II) and thereby alter transcription rates in neurons and other rapid stimulus-induced rapid changes in transcription programs.
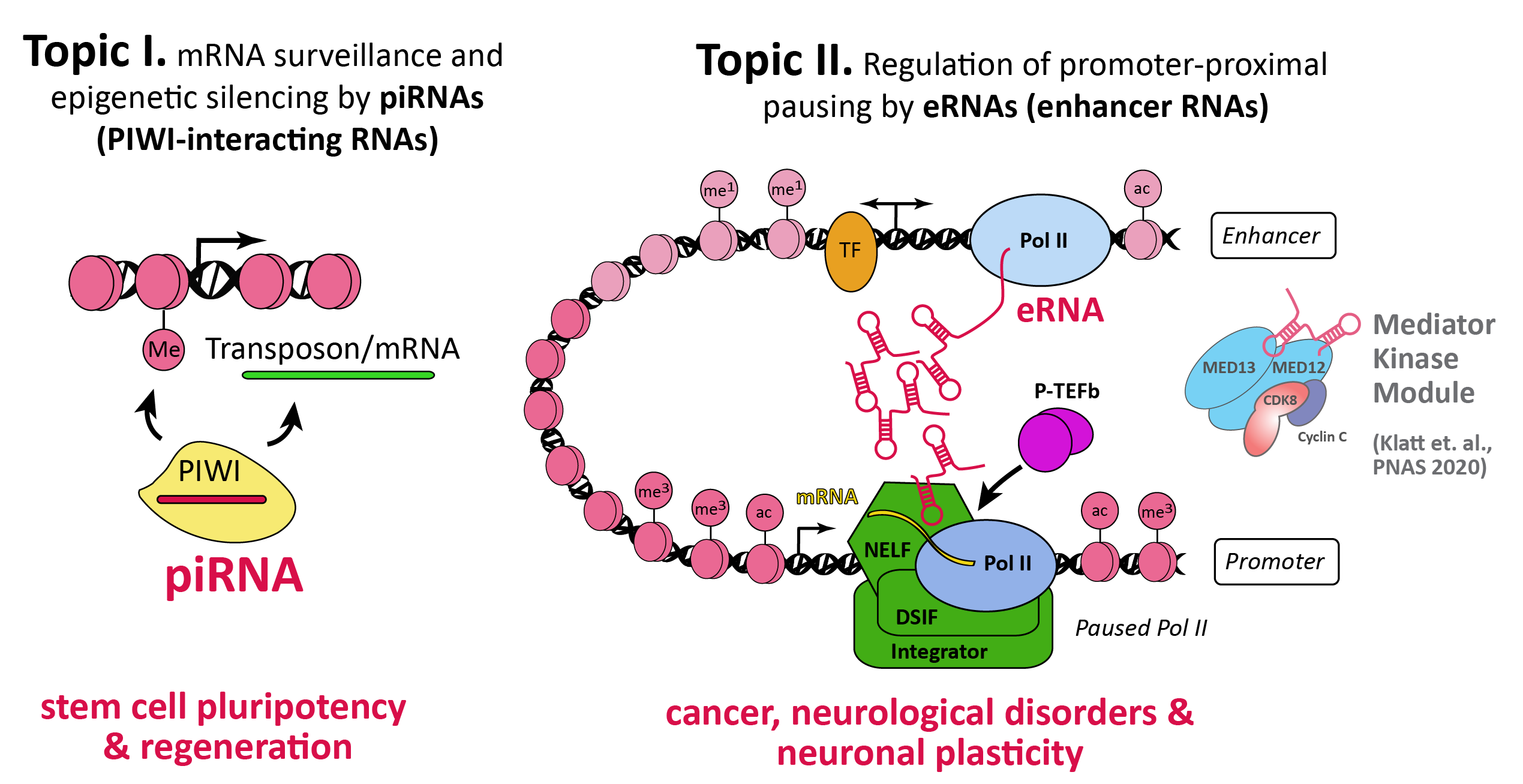
Graphical Summary of Research. The Kuhn lab investigates the essential role of piRNAs (PIWI-interacting RNAs) in the stem cells of highly regeneration-competent planarian flatworms. Specifically, we aim to decipher by how piRNAs direct stem cell mRNA surveillance post-transcriptionally and epigenetically. Second, we examine the mechanistic role that enhancer RNAs (eRNAs) play in neuronal transcription induction by abrogating promoter-proximal pausing. (Our work on the Mediator kinase module (Klatt et al., PNAS 2020) is currently on hold to more quickly advance the other two topics in the lab)

